This was a project performed for a data science degree through the Universities of Wisconsin.
This project created a controlled software environment for potential use in FDA-regulated research, development, and manufacturing. Biopython was used within the environment to analyze protein sequence data from NCBI to identify potential target sequences for recombinant manufacturing in E. Coli.
The environment was built using Docker, which included Git, AWSCLI, and Micromamba to manage Python dependencies for Biopython, boto3, Jupyter, and pyMSAviz. Compliance features (user access control, version history, and controlled storage) were included with Git/GitHub and AWS. The project structure is shown below.
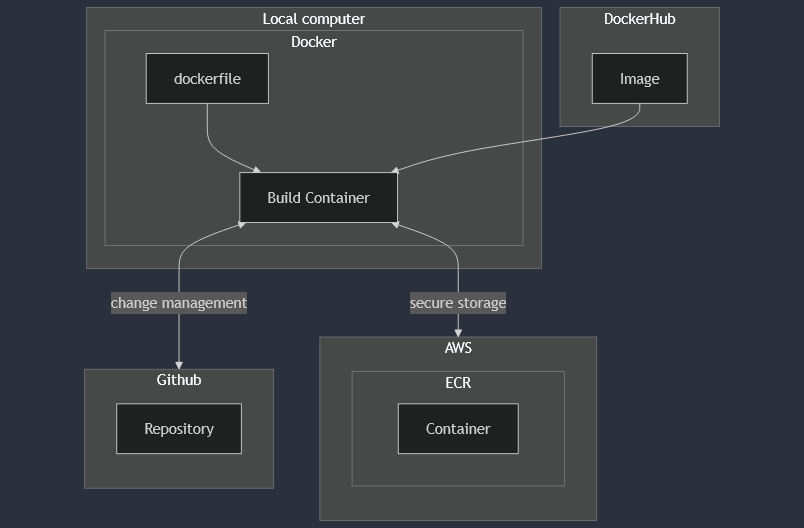
The container was deployed on a local machine for protein sequence analysis of insulin. Data was sourced from NCBI using their BLAST tool to extract FASTA files, then cleaned and filtered for alignment in the software MEGA. The aligned sequences were annotated, analyzed, and compared for desired characteristics to find the most promising targets for future development. How the container was utilized is shown below.
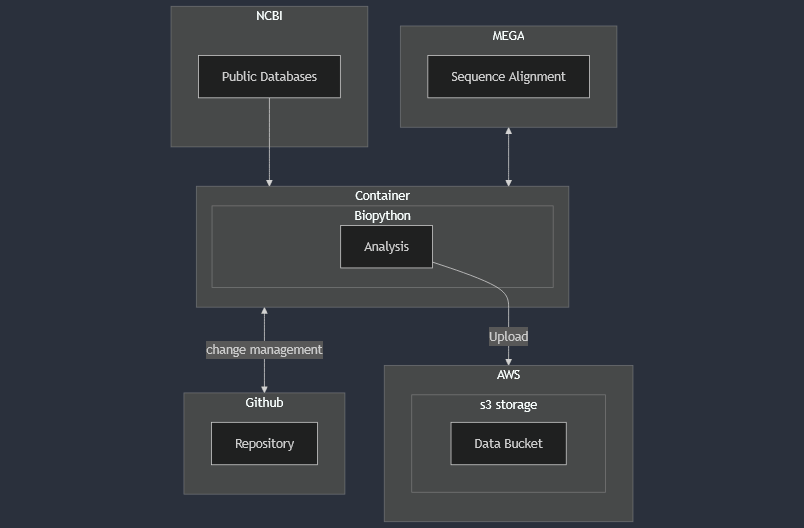
The example analysis done for this project was on insulin, which is primarily recombinantly produced in bacteria (E. Coli) or yeast (S. Cerevsiae) today. Current production methods result in non-functional protein aggregates called inclusion bodies that need further processing to be usable. Inclusion body formation should be avoided where possible, since it limits the ability of microorganisms to continue producing target proteins and adds complexity and cost to manufacturing.
Data analysis was done using Python (within the container), and raw data was sourced from NCBI using their BLAST tool and BLOSUM62 scoring. Protein sequences were aligned with the MEGA program with MSA algorithms ClustalW and Muscle. Aligned sequences were annotated and filtered for those with the least reliance on scarce amino acids (arginine, isoleucine, leucine, proline, and glycine) for E. Coli, and cysteine. Insulin structure and sequence was, in general, highly conserved, especially for the functional A and B chains.
To see if the resulting target sequences were reasonable, they were compared to human insulin (Homo Sapiens) with a phylogenetic tree, shown below. The results are promising because they are primarily derived from mammals. The synthetic sequence included in the data was developed for thermo-induced production in E. Coli – the same use case this analysis attempted to optimize for.
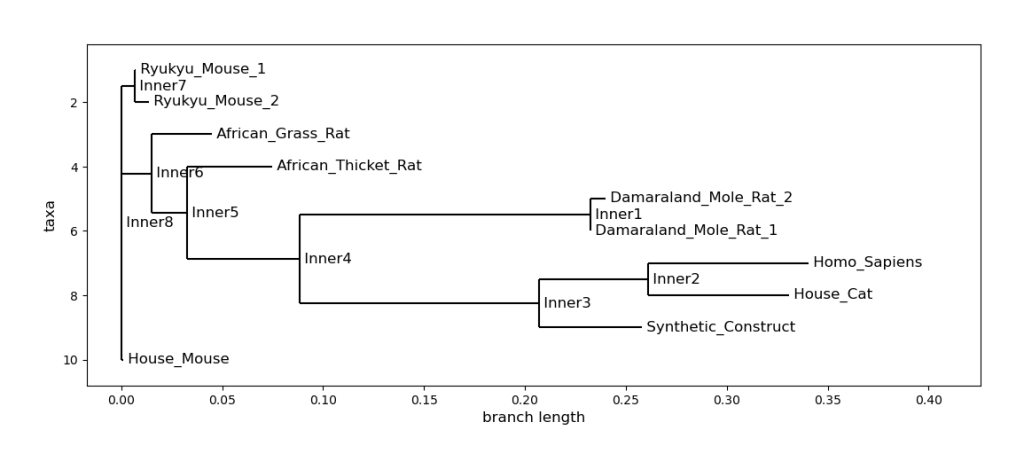
Leave a comment